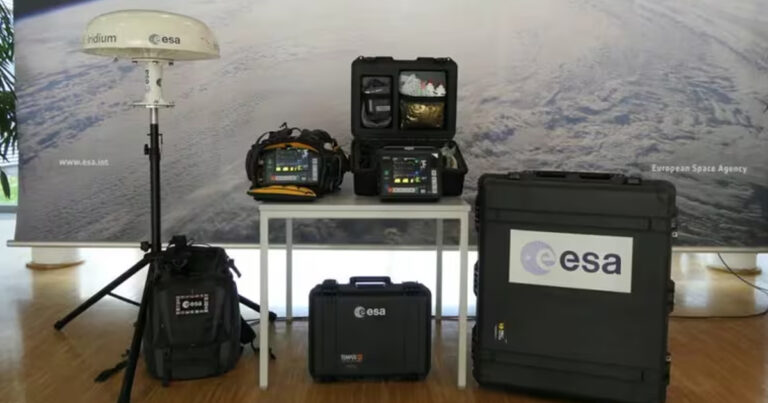
El Valor de la Salud Digital
Es el catalizador de la transformación de los sistemas de salud, a fin de mejorar el acceso, la cobertura efectiva a servicios de salud de calidad, eficientes y con sentido humano, que mejoren la vida de las personas.

Educación Profesional Continua
Plataforma de capacitación más grande de América Latina. Incluye Contenidos Actualizados y curados sobre COVID-19, así como cursos prácticos de diversos temas relevantes. Dirigido a la población en general y para profesionales de la salud.
CLIKISalud
Información y materiales relacionados para el cuidado de tu salud y la de tu familia, para todas las edades.
CLIKISalud cuenta con información de 21 temas en salud, con tips prácticos para implementar en tu vida cotidiana.